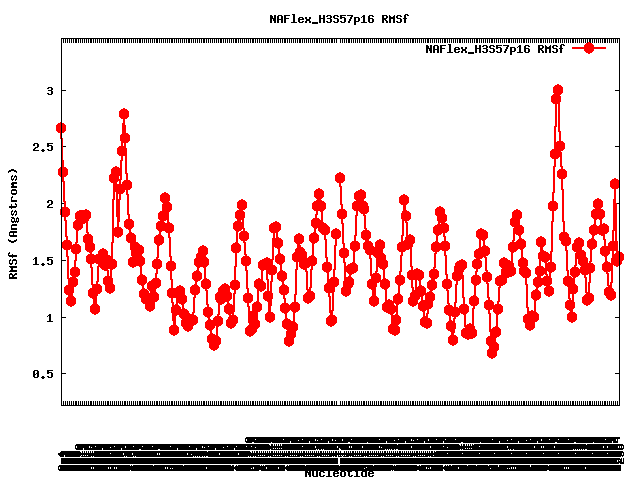
Entry: NAFlex_H3S57p16
Nucleic Acid Data:
| Sequence 1: | ATCAATATCCACCTGCAGATACTACCAAAAGTGTATTTGGAAACTGCTCCATCAAAAGGCATGTTCAGCTGGAATCCAGCTGAACATGCCTTTTGATGGAGCAGTTTCCAAATACACTTTTGGTAGTATCTGCAGGTGGATATTGAT |
| Sequence 2: | ATCAATATCCACCTGCAGATACTACCAAAAGTGTATTTGGAAACTGCTCCATCAAAAGGCATGTTCAGCTGGATTCCAGCTGAACATGCCTTTTGATGGAGCAGTTTCCAAATACACTTTTGGTAGTATCTGCAGGTGGATATTGAT |
| Type | DNA |
| SubType | B |
| Chains | duplex |
| Pdb | 1KX5 [PDB] [NDB] |
| Ligands | Protein |
| Keywords | DNA-B Duplex Naked ParmBSC1 TIP3P AddedSalt |
MD Simulation >> (Click to expand/shrink)
| Simulation Metadata | |
|---|---|
| Force Field | parmBSC1 |
| Simulation Date | 2018-Sep-10 |
| Simulated Time | 700 ns |
| Time Step | 100 ps |
| Parts | DNA, nucleosome core particle |
| Temperature | 300K |
| Water | TIP3P |
| Additional Solvent | No |
| Counter Ions | K+ |
| Ionic Concentration | 0.15 M |
| Additional Ions | KCl |
| Additional Molecules | No |
| Ions Parameters | Dang |
Unified Molecular Modeling (UMM) Metadata
(Click to see full UMM)
(Click to see full UMM)
| Simulation Metadata (UMM) | |
|---|---|
| Author | Pablo Dans |
| AdditionalIons | KCl |
| AdditionalMolecules | No |
| AdditionalSolvent | No |
| Author | . |
| Chains | duplex |
| Comments | 700 ns of MD simulation of a modified H3S57p nucleosome. |
| CounterIons | K+ |
| Format | mdcrd |
| FrameStep | 100 ps |
| Frames | 7000 |
| IonicConcentration | 0.15 M |
| IonsParameters | Dang |
| Ligands | Protein |
| NucType | DNA |
| PDB | 1KX5 |
| Parts | DNA, nucleosome core particle |
| RMSd_avg | 2.187 |
| RMSd_stdev | 0.250 |
| Rgyr_avg | 45.042 |
| Rgyr_stdev | 0.549 |
| SASA_avg | 51013.972 |
| SASA_stdev | 622.825 |
| SubType | B |
| Temperature | 300K |
| Topology | H3S57p_dry.top |
| Trajectory | H3S57p_dry.trj.gz |
| Water | TIP3P |
| _id | NAFlex_H3S57p16 |
| dataset | Array |
| date | 2018-Sep-10 |
| description | DNA-B Duplex Naked ParmBSC1 TIP3P AddedSalt |
| forceField | parmBSC1 |
| ionsModel | - |
| moleculeType | Dna |
| rev_sequence | XX |
| saltConcentration | - |
| sequence | ATCAATATCCACCTGCAGATACTACCAAAAGTGTATTTGGAAACTGCTCCATCAAAAGGCATGTTCAGCTGGAATCCAGCTGAACATGCCTTTTGATGGAGCAGTTTCCAAATACACTTTTGGTAGTATCTGCAGGTGGATATTGAT | ATCAATATCCACCTGCAGATACTACCAAAAGTGTATTTGGAAACTGCTCCATCAAAAGGCATGTTCAGCTGGATTCCAGCTGAACATGCCTTTTGATGGAGCAGTTTCCAAATACACTTTTGGTAGTATCTGCAGGTGGATATTGAT |
| sequenceMulti | Array |
| soluteAtoms | 9374 |
| soluteCharge | -296 |
| soluteResidues | 296 |
| solventAtoms | - |
| solventResidues | - |
| time | 700 |
| totalAtoms | 9374 |
| totalCharge | -296 |
| totalIons | - |
| totalResidues | 296 |