Entry: NAFlex_Z-DNA-0M
MD Simulation >> (Click to expand/shrink)
| Simulation Metadata | |
|---|---|
| Force Field | parmBSC1 |
| Simulation Date | 2015-Apr-25 |
| Simulated Time | 385 ns |
| Time Step | 2ps |
| Parts | DNA |
| Temperature | 300K |
| Water | TIP3P |
| Additional Solvent | No |
| Counter Ions | Na+ |
| Ionic Concentration | Electroneutrality |
| Additional Ions | No |
| Additional Molecules | No |
| Ions Parameters | Dang |
Unified Molecular Modeling (UMM) Metadata
(Click to see full UMM)
(Click to see full UMM)
| Simulation Metadata (UMM) | |
|---|---|
| AdditionalIons | No |
| AdditionalMolecules | No |
| AdditionalSolvent | No |
| Author | I.I. |
| Chains | duplex |
| Comments | - |
| CounterIons | Na+ |
| Format | mdcrd |
| FrameStep | 2ps |
| Frames | 192787 |
| IonicConcentration | Electroneutrality |
| IonsParameters | Dang |
| Ligands | No |
| NucType | DNA |
| PDB | 1I0T |
| Parts | DNA |
| RMSd_avg | 2.973 |
| RMSd_stdev | 0.368 |
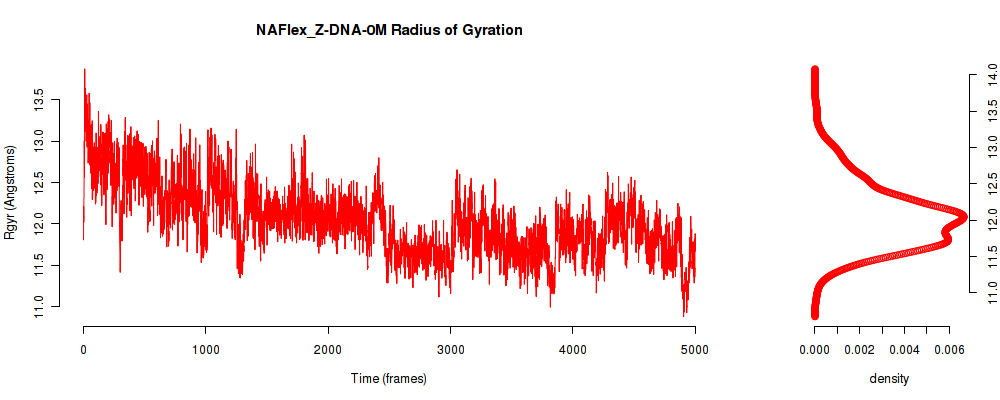
| Rgyr_avg | 12.040 |
| Rgyr_stdev | 0.459 |
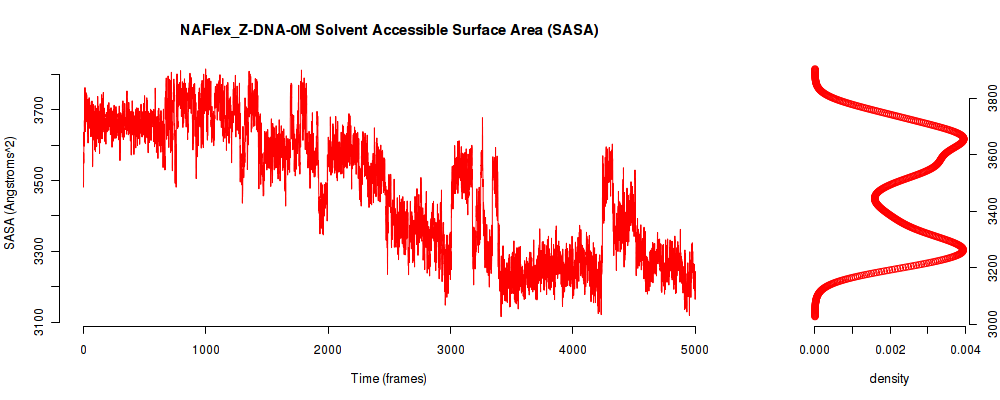
| SASA_avg | 3473.717 |
| SASA_stdev | 187.159 |
| SubType | Z |
| Temperature | 300K |
| Topology | no_wat.top |
| Trajectory | zdna0M/naked.trj.gz |
| Water | TIP3P |
| _id | NAFlex_Z-DNA-0M |
| altPDB | 131D 133D 1D24 1D33 1D39 1D48 1DA2 1DCG 1DJ6 1DN4 1DN5 1G3A 1I0T 1I7J 1ICK 1PBL 1PBM 1QDA 1T4X 1TNE 1UQG 1V9G 1VRO 1VTU 1VTV 1WOE 223D 242D 277D 292D 293D 2DCG 2ELG 2IE1 310D 336D 390D 391D 392D 3P4J 3WBO 427D 4FS5 4FS6 4HIF 4HIG 4R15 |
| authors | Ivan Ivani, Pablo D. Dans, Agnès Noy, Alberto Pérez, Ignacio Faustino, Adam Hospital, Jürgen Walther, Pau Andrio, Ramon Goñi, Alexandra Balaceanu, Guillem Portella, Federica Battistini, Josep Lluís Gelpí, Carlos González, Michele Vendruscolo, Charles A. Laughton, Sarah A. Harris, David A. Case and Modesto Orozco |
| dataset | Array |
| date | 2015-Apr-25 |
| description | DNA-Z Duplex Naked ParmBSC1 TIP3P Electroneutral |
| forceField | parmBSC1 |
| ionsModel | - |
| moleculeType | Dna |
| rev_sequence | CGCGCGCGCG |
| saltConcentration | - |
| sequence | CGCGCGCGCG |
| soluteAtoms | 628 |
| soluteCharge | -18 |
| soluteResidues | 20 |
| solventAtoms | - |
| solventResidues | - |
| time | 385 |
| totalAtoms | 628 |
| totalCharge | -18 |
| totalIons | - |
| totalResidues | 20 |