Entry: NAFlex_groovebinder
MD Simulation >> (Click to expand/shrink)
| Simulation Metadata | |
|---|---|
| Force Field | parmBSC1 |
| Simulation Date | 1429920000000 |
| Simulated Time | 700 |
| Time Step | 100ps |
| Parts | only DNA |
| Temperature | 300K |
| Water | TIP3P |
| Additional Solvent | No |
| Counter Ions | Na+ |
| Ionic Concentration | Electroneutrality |
| Additional Ions | No |
| Additional Molecules | No |
| Ions Parameters | Dang |
Unified Molecular Modeling (UMM) Metadata
(Click to see full UMM)
(Click to see full UMM)
| Simulation Metadata (UMM) | |
|---|---|
| AdditionalIons | No |
| AdditionalMolecules | No |
| AdditionalSolvent | No |
| Author | I.F. |
| Chains | duplex |
| Comments | - |
| CounterIons | Na+ |
| Format | mdcrd |
| FrameStep | 100ps |
| Frames | 7000 |
| IonicConcentration | Electroneutrality |
| IonsParameters | Dang |
| Ligands | No |
| NucType | DNA |
| PDB | 2DND |
| Parts | only DNA |
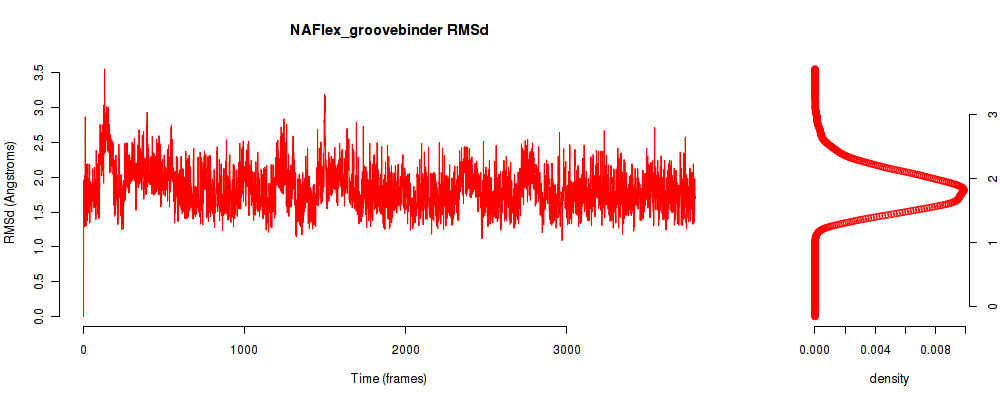
| RMSd_avg | 1.840 |
| RMSd_stdev | 0.304 |
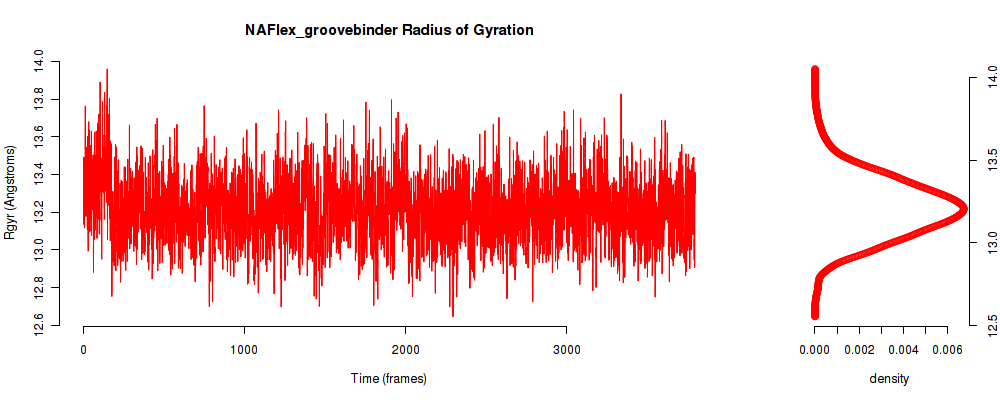
| Rgyr_avg | 13.209 |
| Rgyr_stdev | 0.278 |
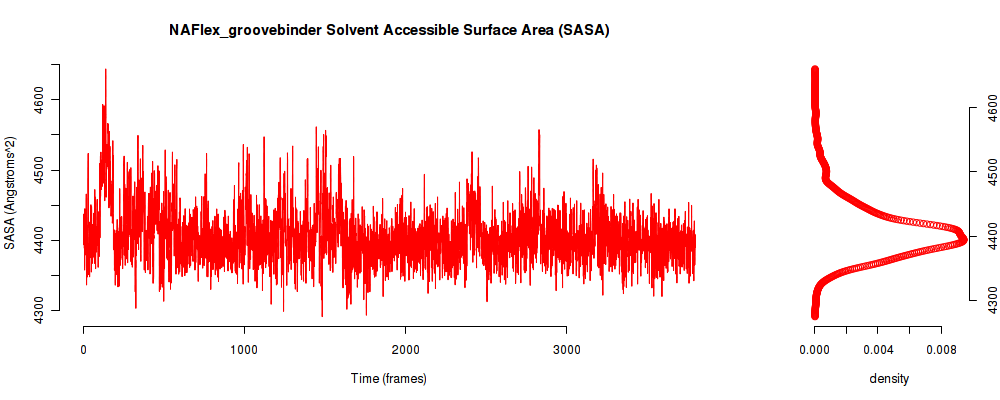
| SASA_avg | 4404.680 |
| SASA_stdev | 81.621 |
| SubType | B |
| Temperature | 300K |
| Topology | groovebinder.strip.top |
| Trajectory | groovebinder.onlydna.trj.gz |
| Water | TIP3P |
| _id | NAFlex_groovebinder |
| altPDB | 102D 121D 1D63 1D65 1DXN 1L3M 1S2R 263D 264D 296D 2DND 4AH0 4AH1 |
| authors | Ivan Ivani, Pablo D. Dans, Agnès Noy, Alberto Pérez, Ignacio Faustino, Adam Hospital, Jürgen Walther, Pau Andrio, Ramon Goñi, Alexandra Balaceanu, Guillem Portella, Federica Battistini, Josep Lluís Gelpí, Carlos González, Michele Vendruscolo, Charles A. Laughton, Sarah A. Harris, David A. Case and Modesto Orozco |
| dataset | Array |
| date | 1429920000000 |
| description | DNA-B Duplex Naked ParmBSC1 TIP3P Electroneutral minor-groove-binder |
| forceField | parmBSC1 |
| ionsModel | - |
| moleculeType | Dna |
| rev_sequence | CGCAAATTTGCG |
| saltConcentration | - |
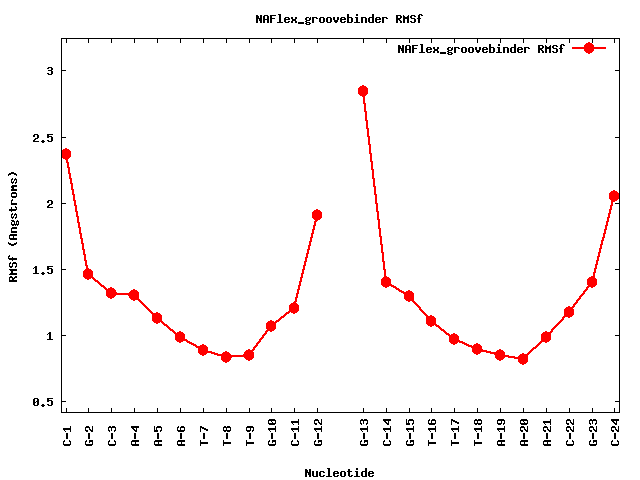
| sequence | CGCAAATTTGCG |
| soluteAtoms | 760 |
| soluteCharge | -22 |
| soluteResidues | 24 |
| solventAtoms | - |
| solventResidues | - |
| time | 700 |
| totalAtoms | 760 |
| totalCharge | -22 |
| totalIons | - |
| totalResidues | 24 |