Entry: NAFlex_1g14
MD Simulation >> (Click to expand/shrink)
| Simulation Metadata | |
|---|---|
| Force Field | parmBSC1 |
| Simulation Date | 1437091200000 |
| Simulated Time | 1,000 |
| Time Step | 200 ps |
| Parts | DNA |
| Temperature | 298K |
| Water | TIP3P |
| Additional Solvent | No |
| Counter Ions | Na+Cl- |
| Ionic Concentration | 0.15M |
| Additional Ions | No |
| Additional Molecules | No |
| Ions Parameters | Dang |
Unified Molecular Modeling (UMM) Metadata
(Click to see full UMM)
(Click to see full UMM)
| Simulation Metadata (UMM) | |
|---|---|
| AdditionalIons | No |
| AdditionalMolecules | No |
| AdditionalSolvent | No |
| Author | G.P. |
| Chains | duplex |
| Comments | - |
| CounterIons | Na+Cl- |
| Format | xtc |
| FrameStep | 200 ps |
| Frames | 5000 |
| IonicConcentration | 0.15M |
| IonsParameters | Dang |
| Ligands | No |
| NucType | DNA |
| PDB | 1G14 |
| Parts | DNA |
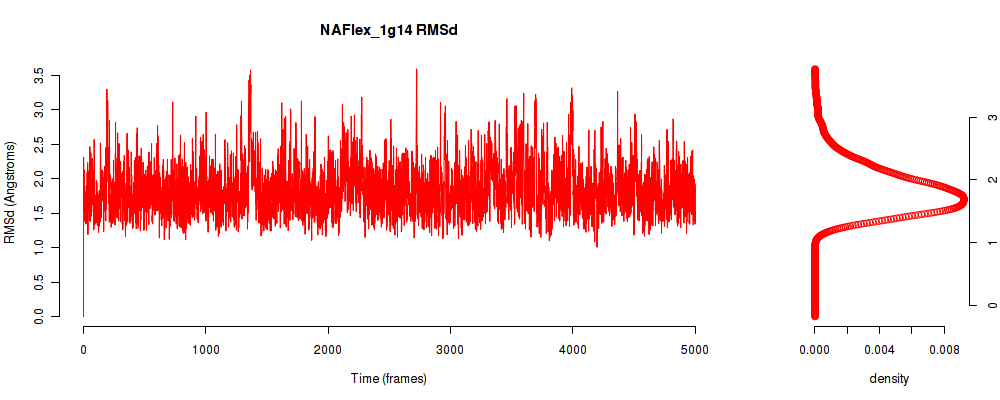
| RMSd_avg | 1.835 |
| RMSd_stdev | 0.363 |
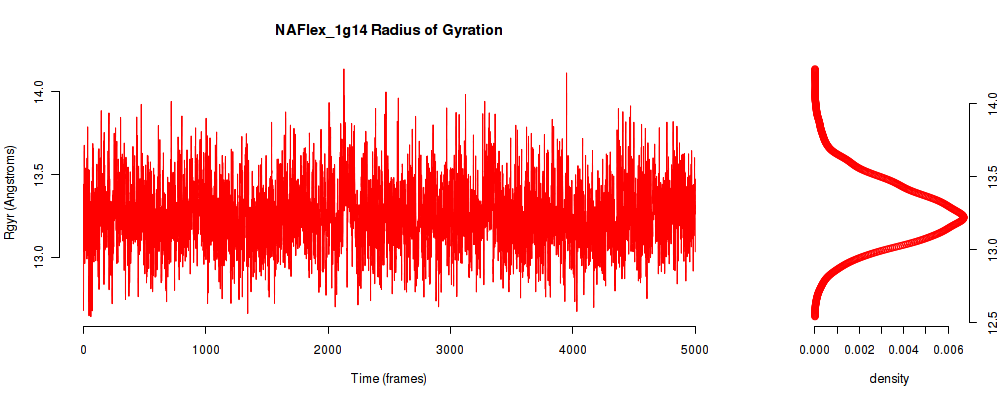
| Rgyr_avg | 13.251 |
| Rgyr_stdev | 0.279 |
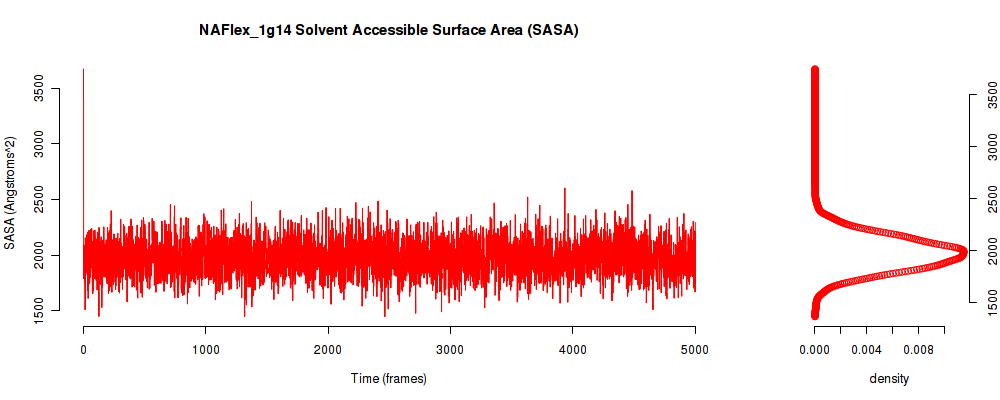
| SASA_avg | 1975.732 |
| SASA_stdev | 161.543 |
| SubType | B |
| Temperature | 298K |
| Topology | init_1g14.pdb |
| Trajectory | 1g14_bsc1_5000.xtc |
| Water | TIP3P |
| _id | NAFlex_1g14 |
| dataset | Array |
| date | 1437091200000 |
| description | DNA-B Duplex Naked Parm99 TIP3P AddedSalt |
| forceField | parm99 |
| ionsModel | - |
| moleculeType | Dna |
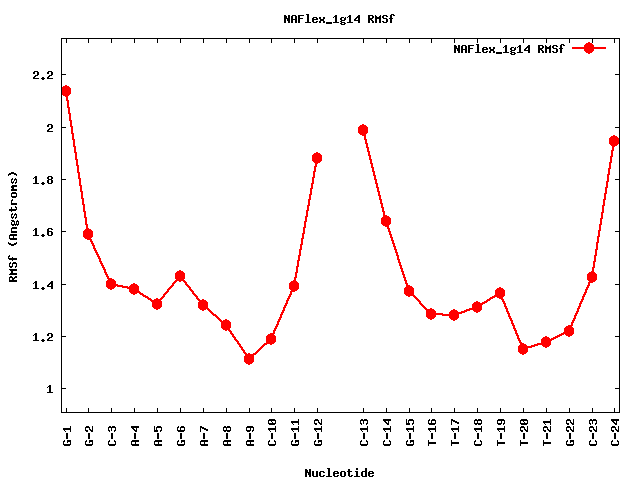
| rev_sequence | CCGTTTCTTGCC |
| saltConcentration | - |
| sequence | GGCAAGAAACGG |
| soluteAtoms | 759 |
| soluteCharge | 759 |
| soluteResidues | 24 |
| solventAtoms | - |
| solventResidues | - |
| time | 1,000 |
| totalAtoms | 759 |
| totalCharge | 759 |
| totalIons | - |
| totalResidues | 24 |