BIGNASim database structure and analysis portal for nucleic acids simulation data
Database Structure
BigNaSim is based on the combination of two database engines, Cassandra and MongoDB, and an adapted version of the analysis section of our Nucleic Acids MD portal, NAFlex (1). For trajectory data manipulation, the platform uses MDPlus, an in-house python library that integrates MDAnalysis tools (2) with a developed Cassandra interface.
Cassandra. The trajectory subsystem.
Cassandra (3) is a distributed and highly scalable key-value database with a strong user community. Cassandra implements a non-centralized architecture, based on peer-to-peer communication, in which all nodes of the cluster are able to receive and serve queries. Data is stored in tables by rows, which are identified by a key chosen by the database user. In each row, users can add different attributes also identified by a chosen name. To each node of the cluster a token is assigned and become responsible for hosting specific set of rows. The target node for each row is chosen through the partitioner algorithm and the decision is based on the row key and the node token. Cassandra also allows using compound keys and thus, more than one attributes to identify a row. In this case, the users have to specify the partition key (i.e. the attribute that will guide the node assignment) and the clustering keys. When a node receives a query, it finds out which node is responsible for each row involved in the query and partitions, and forwards appropriately the data request. This means that data modelling has a key influence on query performance (4). For this reason, users are encouraged to define their data models considering which queries they are going to perform. Moreover, a common practice is to replicate data in different data models to accommodate different queries. Recently, the authors proposed a mechanism to alleviate users from this requirement (5).
Table S2. Cassandra trajectory database structure.
| Topology table (idSimulation) | Trajectory table (idSimulation) |
| atom_num (Partition Key) atom_name atom_type chain_code residue_code residue_num |
frame (Partition Key) atom_id (Clustering Key) x y z (Box size data is included in the same frame as additional pseudo-atoms) |
The Cassandra subsystem was organized in two tables: Topology holds the description of the molecular system using atom number as main indexing key, and storing the atom details, and the usual logical ways of grouping them (residue, chain). The Trajectory table stores the coordinates themselves indexed using frame and atom numbers. Cassandra is a distributed system, and the selection of the partition key has a strong influence on the retrieval efficiency. In our implementation trajectory data is distributed using frame numbers, improving the retrieval of frame blocks. Indeed, by defining the frame number as Partition Key, we ensure that all the atomic coordinates at a given snapshot are stored contingently in the same node. Additionally, each frame has atomic identifiers as a second level index, allowing efficient access to any subset of atoms. We have chosen to prioritize frame-based access, after analyzing the pattern of access of the MDAnalysis software, used to handle trajectory data. MDAnalysis, constrained by its interface, always access to trajectory a frame at a time. Consequently, with our model the existing algorithms can access to a trajectory in Cassandra seamless, as if it was a common file. At the same time, algorithms that require data of only a subset of atoms, may be optimized to take advantage of the second level indexing. To move trajectory data in and out of the Cassandra subsystem, the use of the Python package MDPlus assures a full compatibility with existing molecular dynamics software. Still, when dealing with massive bulk data loading into the database, the overhead introduced by the network communications and the data marshalling between different platforms can be a problem. For that reason, we developed a utility program that takes as input a trajectory file and converts it directly into SSTables, the Cassandra internal data format.
MongoDB. The analysis and metadata subsystem
The MongoDB database holds simulation metadata and pre-calculated analysis results. MongoDB is a fully flexibly engine and can store heterogeneous collections of documents. The internal structure of each document does not need to be defined beforehand and can match the data structure used in the interacting software, thus simplifying the use of database documents and external analysis software. MongoDB also allows to partition data among different servers (data sharding), using any of the fields as partition key. In our case, the data of the analysis requires both frame-based and atom-based access, hence we have chosen the complete document key as sharding key (See Table S3 below). Although MongoDB is configured with a single entry-point, it processes access queries in parallel among the available nodes, so maximum efficiency is achieved when data is spread evenly among them.
A condition to make the database usable is a very consistent indexing schema, which allows an easy recovery of such documents. Table S3 shows the database collection list together with the primary keys used to store the different objects. Table S4 shows representative data objects as stored in the DB.
Table S3. Structure of main MongoDB collections
| MongoDB collection | Main Index components | Description |
| simData | idSim | Simulation metadata, following a specifically defined ontology |
| analDefs | idSim, idAnal | Analysis description, one document stored for every analysis result item available. Analysis available could differ from one simulation to another |
| groupDef | IdSim, (idGroup,nGroup) | Molecular groups (bases, base pairs, base-pair steps, molecular fragments) defined in the simulated system |
| analData | idSim, (idGroup,nGroup), nFrame
(nFrame = 0: Averaged analysis data) |
Analysis results. The most appropriate data model for each analysis type is used. |
| analBinFiles | Id. Above | Binary files with pre-calculated analysis results (plots, images, etc.) |
Fragment definition
MongoDB BIGNASim database has been populated using in-house scripts, and parsing the results obtained from the series of well-known software of analysis implemented in NAFlex (1). Definition of residues and standard groups (nucleotides, base-pair, base-pair steps) are generated automatically from the simulated sequence and stored in the groupDef collection. Besides of predefined standard groups, the collection can store the definition of any relevant fragment of the simulated molecular system. As a representative example or groupDef structure, Figure S2 shows the complete hierarchy derived from the central tetramer of a Drew-Dickerson dodecamer.
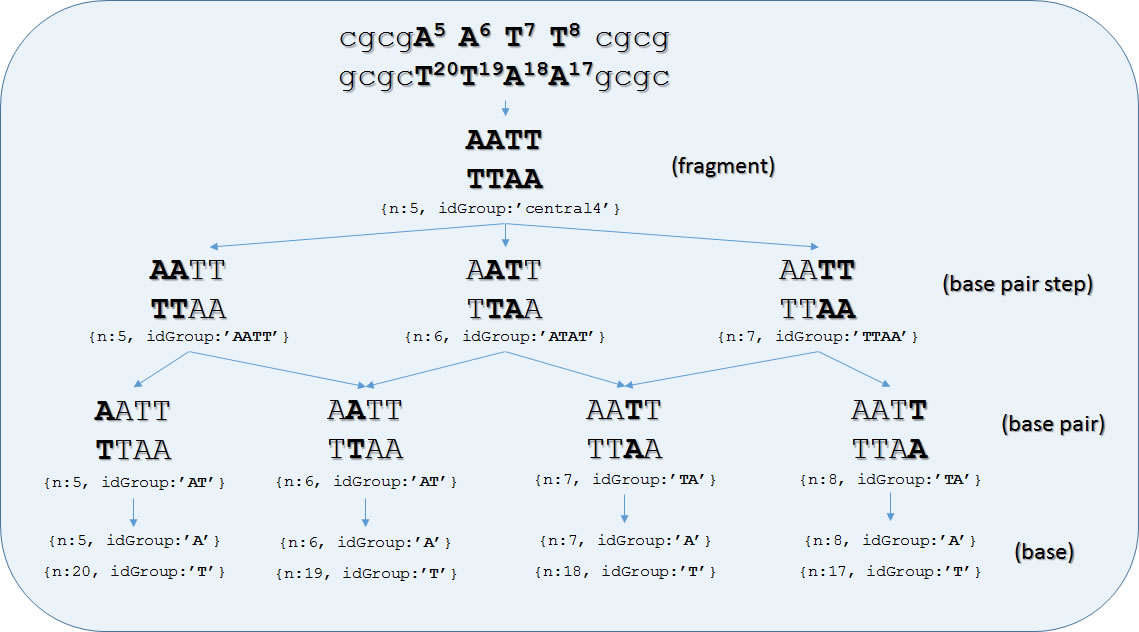
Figure S2. Example of fragment definition on the analysis database. Database entries in groupDef collection derived from the central tetramer of a Drew-Dickerson dodecamer. Primary keys of each data item are indicated. Arrows indicate a “container” relationship between data objects. Simulation id has been deleted from keys for simplicity.
The complete structure of such objects can be found in Table S5. Once the fragments are defined, their id (composed by idSim, idGroup and nGroup, see Table S3) are used to index analysis results in analData and analBinFiles collections. As shown in Figure S2 above, and Table S5, the collection also holds a hierarchic relationships indicating which are the components of each fragment from the immediate lower level; this allows to navigate from any group down to its composing parts, and to the individual bases (see use case 4, for an example of such usage). This would allow linking together analysis corresponding to the related hierarchical levels. At the residue level, the MongoDB analysis subsystem is consistent with data hold in the trajectory subsystem (i.e. idGroup + nGroup corresponds to residue_code + residue_num). As shown in Table S2, results of the analyses are again stored in the three axes space: simulation, the analysed group (split in group id and sequence number for convenience), and frame number. Also, averages along the trajectory and analysis spanning the whole system can be stored in the same structure. This layout will allow retrieving easily any set of results for any given set of groups and frames and performing the appropriate post-process. Although most data can be retrieved from the BIGNASim portal, more specialized data combination would require specific scripts. See examples of use for examples of such scripts using MongoDB JavaScript. Similar scripts can be prepared using other programming languages (Python, Perl, PHP, Java). Analyses that may lead to non-numerical results (XY plots, 3D grids, etc.) are also stored under the same coordinate system, although they are kept in a separate collection (analBinFiles) for efficiency reasons. This database layout could be extended to any new type of analysis, without modification, after an appropriate mapping of each individual data item in the group/frame axes. Additionally, the GridFS system provided by MongoDB has been used to handle file based data transfers between application modules, and to hold the temporary user space used for downloading data.

