Browsing
All nucleic acids MD simulation data stored in our system can be accessed from the BIGNASim portal browse page.
Simulations are shown in a table together with a short description including essential information such as nucleic acid type (DNA/RNA/Complex/etc.), subtype (single/duplex/etc.), and essential MD parameters such as force-field or solvent type used. Also, simulation length, and nucleotidic sequence are shown (see figure below). Simulations appearing on the screen can be ordered by any of these parameters.
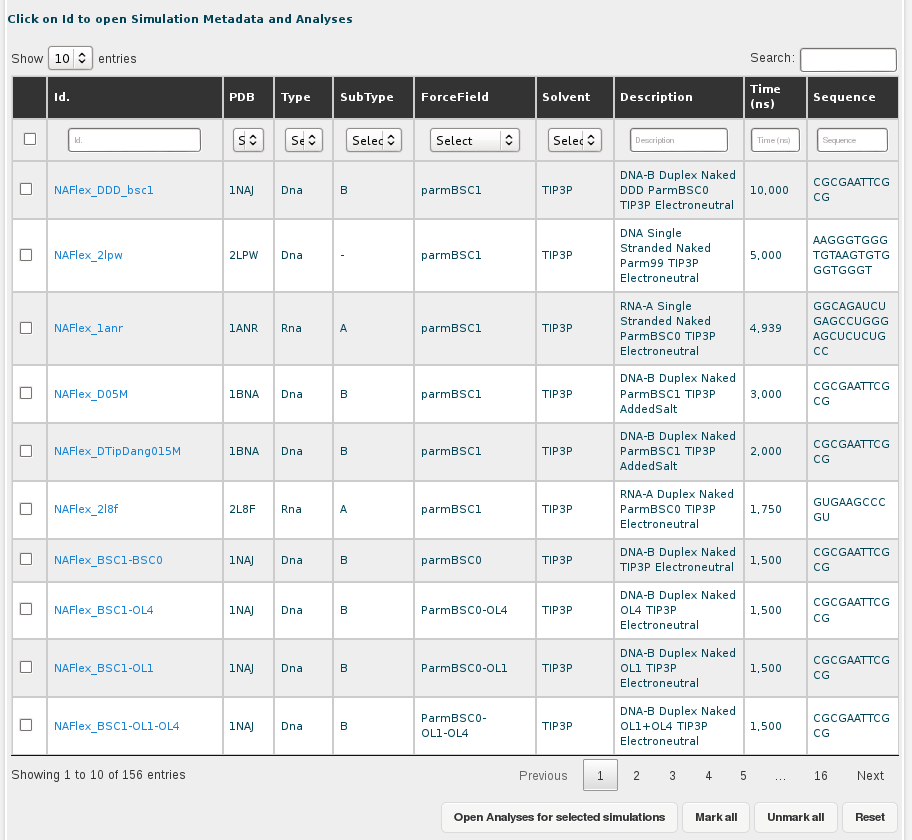
With this interface, the whole database can be easily browsed, and interesting simulations can be selected with checkboxes to obtain the pre-computed analyses stored (Retrieve selected button).
The number of simulations shown can be also controlled by a selector on top of the page.
Browsing the whole database is an easy way to have a first contact with the power of the BIGNASim portal. However, if you are interested in a particular nucleotidic sequence or nucleic acid type, or in a particular base-pair step, you should go to the Search section.
Clicking at one of the trajectories the portal will show all the information regarding to its sequence, simulation and trajectory analyses in a new window (see simulation section).