BIGNASim database structure and analysis portal for nucleic acids simulation data
----
DNA & RNA Structure and Helical Parameters Analyses (NAFlex)
----Tutorial 2 -- Global analysis (XCGY)
Tutorial 3 -- Meta-trajectory (XCGY)
Tutorial 4 -- Experimental vs MD analysis
----Statistics
BIGNASim statistics section shows information about the content of the database at this particular moment. Results shown are generated on-the-fly, thanks to the technology used. Examples of information shown are number of trajectories stored, distribution of trajectories by type (DNA, RNA, Complex) or number of bases/base-pairs/base-pair steps analysed.
Statistics are shown in a graphical way through a set of circular diagrams, accompanied by their corresponding values in tables. By default, just the description of the different statistics analyzed are shown, with plots and tables initially compressed. Information inside can be represented just clicking on the description.

Statistics shown are divided in three different sections:
General Statistics
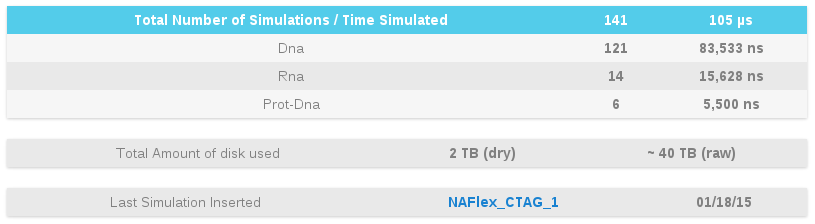
Global statistics about the number of simulations stored, with the corresponding total simulation time represented, or total amount of disk used, are queried on-the-fly and shown in a table.
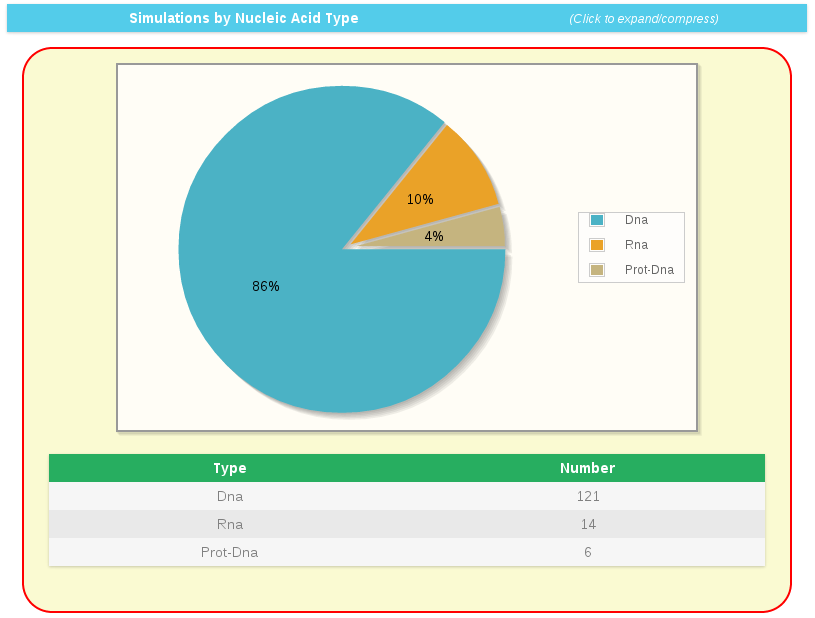
Distribution of simulations stored by nucleic acid type (DNA, RNA, Prot-DNA) is also shown in general statistics. This type of circular plot is used for the rest of presented statistics included in this section of BIGNASim portal.
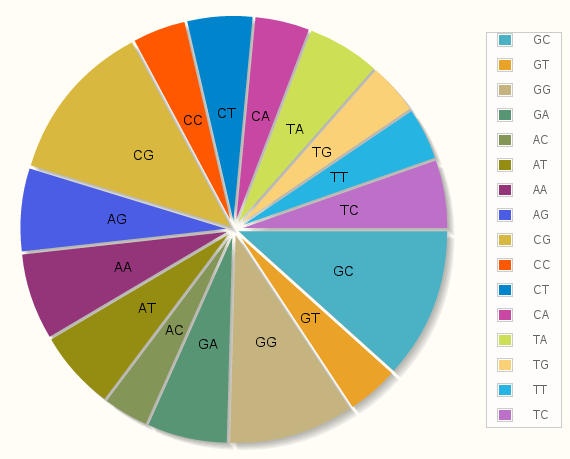
Statistics by Bases / Base-Pairs / Base-Pair Steps
This statistics section shows the representation of different groups of interest building the simulated nucleic acid systems in our database. These groups are bases, base-pairs and base-pair steps.

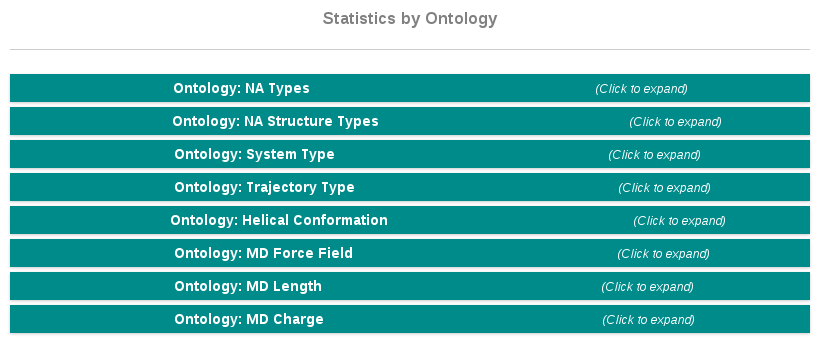
Statistics by Ontology
The nucleic acids ontology generated in this project (see Ontology section) is used not only to classify our systems, but also to build a powerful search engine (see search section) and also allows us to build a set of statistic plots. Statistics sections are divided according to the main fields of the ontology:
- Nucleic Acid Types
- Nucleic Acid Structure Types
- System Types
- Trajectory Types
- Helical Conformation
- MD Force Field
- MD Length
- MD Charge
Each of the sections includes a circular plot with the distribution found in our database, together with a table showing the corresponding values.



