MC DNA Help - Analysis Circular
Circular
This analysis is available when tool “Circular MC DNA” is chosen. The analysis parameters for circular DNA are Twist, Writhe and Radius of gyration. Twist Tw reflects the number of helical turns ($Tw = \sum_{i=1}^{N - 1} tw_i / 360$; N is the length of the sequence, $tw_i$ is the value for Twist in degree of base-pair step i.) and writhe Wr is the number of times the double helix crosses over on itself (supercoils). The relaxed structure for the circle is defined as the structure with Wr = 0 and twist values are the values of the relaxed twist state. Thus the total linking number $Lk_0$ of the relaxed circle is $Lk_0 = Tw_0$. To induce additional stress the twist value of each base pair step of the circle can be changed which results in new value of Tw. Over- or under-twisting of the relaxed structure results in a different linking number $\Delta Lk = Lk - Lk_0 = Tw - Tw_0$ and thus a different starting structure with $\Delta Lk \ne 0$. $\Delta Lk$ can only take integer numbers. $\Delta Lk = Tw + Wr$ will stay constant throughout the whole simulation, however Tw will change throughout the simulation due to the Monte Carlo moves and Wr becomes non-zero. The values of Tw and Wr of the final structure of the simulation are plotted.
Another parameter to analyze the compactness of the circle is the radius of gyration $R_g$. We define the position $r_i$ of base-pair I as the middle between the C6 and C8 atom. The radius of gyration $R_g$ is then calculated as follows:
\[R_g = \sqrt{ \frac{1}{N} \sum_{i=1}^N (r_i - r_{mean})^2} \]
$r_{mean}$ is the mean position of the base-pairs and N is the total number of base-pairs.
For “Structure Flexibility Analysis”
The values of Twist, Writhe and Radius of gyration are given for the relaxed circular structure.
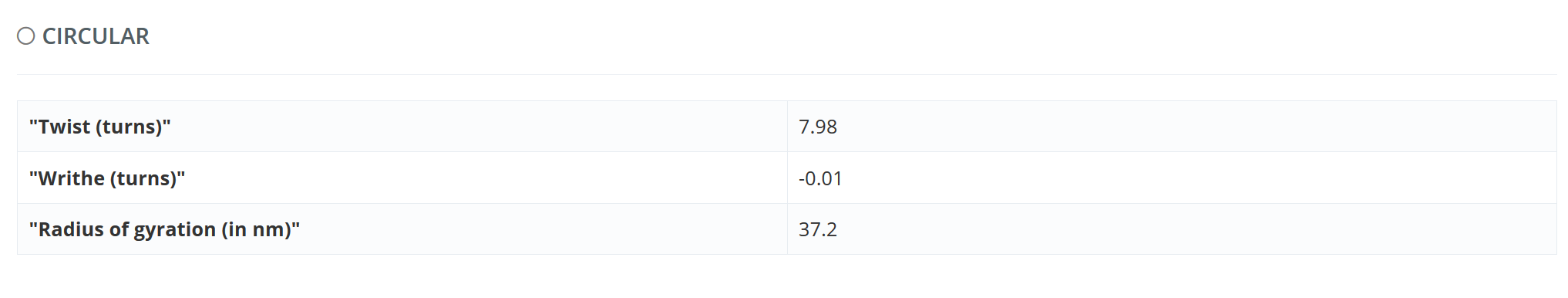
For “Trajectory Flexibility Analysis”
Radius of gyration (in nm), Twist (in turns) and Writhe (in turns) are plotted against the index of the snapshot of the trajectory.
