MC DNA Help - Analysis Elastic Energy
Elastic Energy
The elastic energy $F_{short}(X)$ of the DNA fiber in the current implementation is calculated as follows:
$$F_{short}(X) = K_i \Delta X_i^2$$
$K_i$ is a 6x6 stiffness matrix obtained by inverting the covariance matrix centered at the global minimum i for all unique base-pair steps in all unique tetrameric environments as derived from ABC microsecond-long MD simulations with the parmbsc1 force-field, $\Delta X_i^2$ is the displacement of the given conformation of base pair step parameters ($X_i$).
Elastic energy of DNA: The table shows mean and standard deviation for the elastic energy of the DNA fiber as well as the average elastic energy per base pair
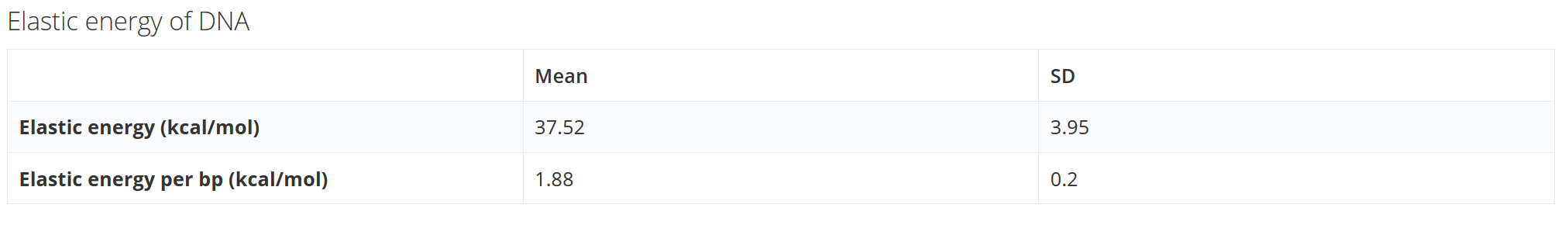
Note that for “Structure Flexibility Analysis” the standard deviation is 0 and the mean represents the elastic energy of the structure.
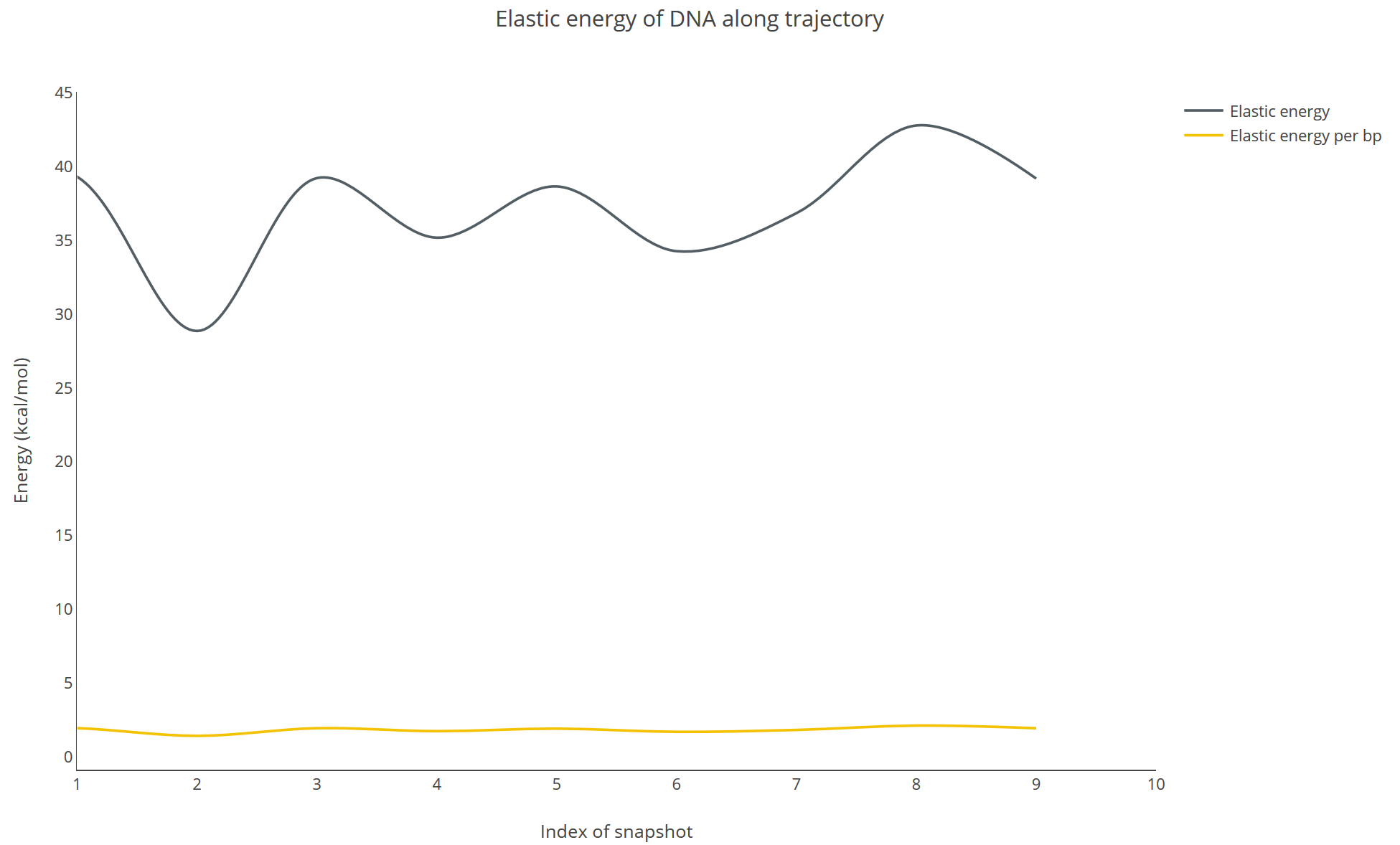
Elastic energy of DNA along trajectory (available for Trajectory Flexibility Analysis)
The elastic energy and the average elastic energy per base pair (y-axis, in kcal/mol) is shown for each snapshot (x-axis) of the trajectory:
Elastic energy of DNA not bound to a protein (available when tool “MC DNA + Proteins” is chosen):
Table with elastic energy and average elastic energy per base pair of the DNA not bound to a protein.
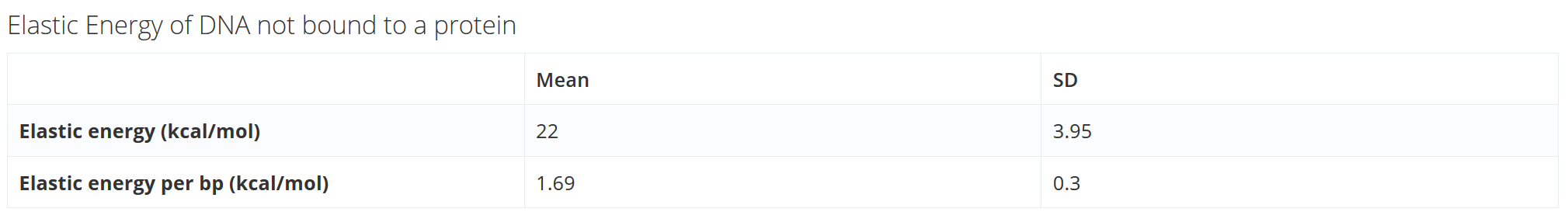
Note that for “Structure Flexibility Analysis” the standard deviation is 0 and the mean represents the elastic energy of the structure.
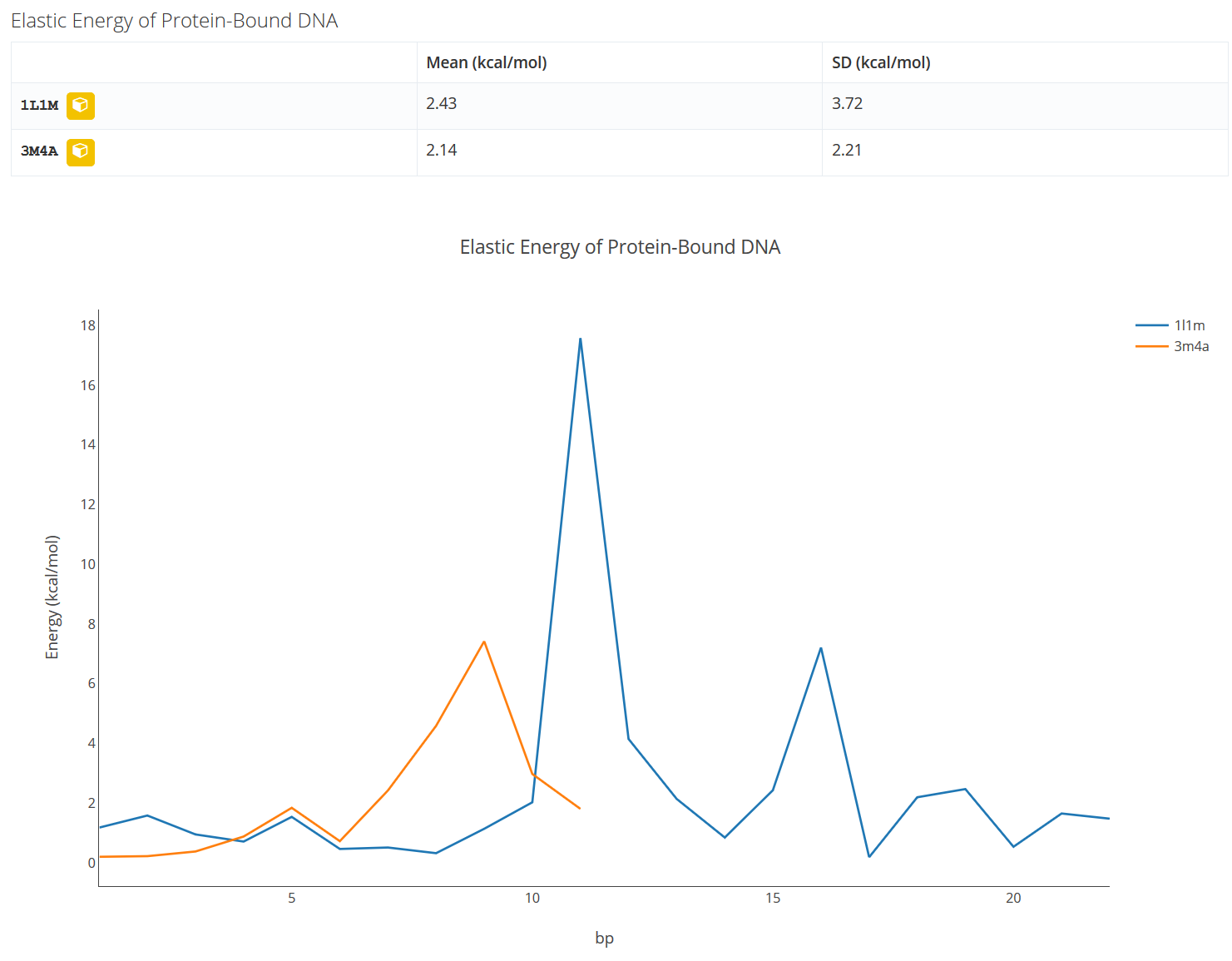
Elastic Energy of Protein-Bound DNA (available when tool “MC DNA + Proteins” is chosen)
A table with the elastic energy of the protein-bound DNA is shown ordered according to the appearance along the DNA fiber. In the plot underneath the values of the elastic energy in kcal/mol (y-axis) along the bound DNA (x-axis) are shown.